2022 Publications
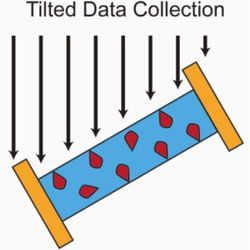
Single-particle cryo-EM data collection with stage tilt using Leginon
A protocol is presented to improve single particle cryo-EM structure determination by tilting specimens while collecting data.
PubMed PMID: 35848829
PubMedCentral PMCID: PMC9733953
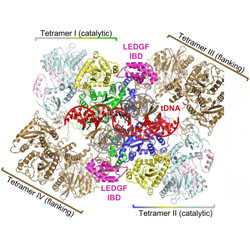
Multivalent interactions essential for lentiviral integrase function
Integrases form multi-subunit intasomes with DNA that may be important for targeting integration to active genes.
PubMed PMID: 35504909
PubMedCentral PMCID: PMC9065133
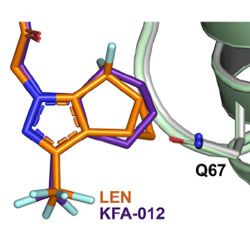
Structural and mechanistic bases of viral resistance to HIV-1 capsid inhibitor Lenacapavir
Structures of two important resistant mutants of HIV-1 capsid allowed the design of a more effective inhibitor.
PubMed PMID: 36190128
PubMedCentral PMCID: PMC9600929
MX2 viral substrate breadth and inhibitory activity are regulated by protein phosphorylation
Phosphorylation sites are identified on MX2 protein that increase HIV-1 inhibition and antiviral activity against mutant viruses.
PubMed PMID: 35880880
PubMedCentral PMCID: PMC9426416
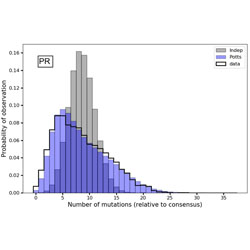
Limits to detecting epistasis in the fitness landscape of HIV
Interactions between mutations affect viral fitness, but may be difficult to observe experimentally.
PubMed PMID: 35041711
PubMedCentral PMCID: PMC8765623
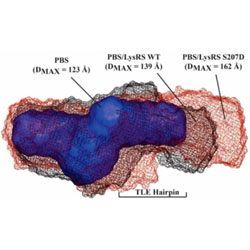
Phosphomimetic S207D lysyl-tRNA synthetase binds HIV-1 5’UTR in an open conformation and increases RNA dynamics
Phosphorylated lysyl-tRNA synthetase may be important for delivering primer tRNAs to HIV-1 genomic RNA.
PubMed PMID: 35891536
PubMedCentral PMCID: PMC9315659
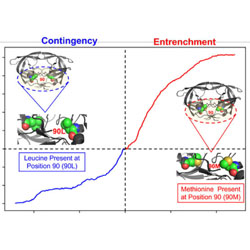
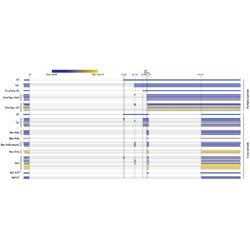
Contingency and entrenchment of drug-resistance mutations in HIV viral proteins
As HIV-1 mutates during antiretroviral therapy, drug resistance mutations are trapped by a combination of many accessory mutations.
PubMed PMID: 36493468
PubMedCentral PMCID: PMC9841799
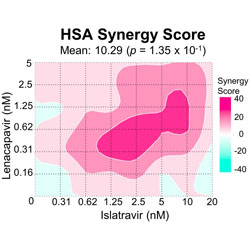
Drug interactions in lenacapavir-based long-acting antiviral combinations
Combinations of capsid inhibitors and reverse transcriptase inhibitors are explored for long-acting treatment of HIV-1.
PubMed PMID: 35746673
PubMedCentral PMCID: PMC9229705
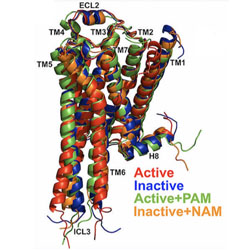
Unique features of different classes of G-protein-coupled receptors revealed from sequence coevolutionary and structural analysis
Covariation analysis of GPCR sequences is used to reveal sites involved in activation and allosteric regulation.
PubMed PMID: 34599827
PubMedCentral PMCID: PMC8738117
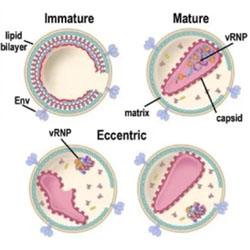
Multimodal functionalities of HIV-1 integrase
B-HIVE Center researchers review HIV-1 integrase mutants that cause defects in maturation.
Engelman AN, Kvaratskhelia M. Viruses. 2022 Apr 28;14(5):926. doi: 10.3390/v14050926.
PubMed PMID: 35632668
PubMedCentral PMCID: PMC9144474
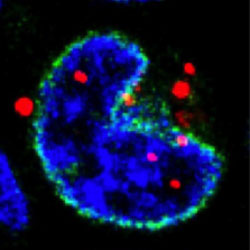
Localization and functions of native and eGFP-tagged capsid proteins in HIV-1 particles
Studies with eGFP-tagged capsid reveal that capsid remodeling is the predominant pathway for HIV-1 nuclear entry.
PubMed PMID: 35951676
PubMedCentral PMCID: PMC9426931
Selective ablation of 3′ RNA ends and processive RTs facilitate direct cDNA sequencing of full-length host cell and viral transcripts
A new sequencing method reveals alternative splicing of RNA within a host cell transcriptome.
PubMed PMID: 35736235
PubMedCentral PMCID: PMC9508845
HIV-1 nucleocapsid protein binds double-stranded DNA in multiple modes to regulate compaction and capsid uncoating
Optical tweezers, fluorescence imaging, and atomic force microscopy are used to explore the interaction of HIV-1 nucleocapsid and DNA.
PubMed PMID: 35215829
PubMedCentral PMCID: PMC8879225

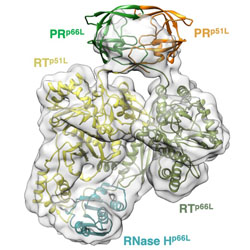
Cryo-EM structure of the HIV-1 Pol polyprotein provides insights into virion maturation
HIV-1 reverse transcriptase forms a similar dimer in the Pol polyprotein and in the mature form.
PubMed PMID: 35857464
PubMedCentral PMCID: PMC9258950
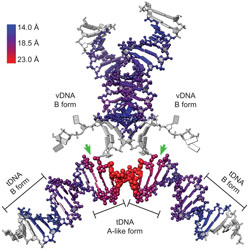
B-to-A transition in target DNA during retroviral integration
Retroviruses selectively integrate into regions of the genome where the host DNA can preferentially adopt the A-form.
PubMedPMID: 35947647
PubMedCentral PMCID: PMC9410886
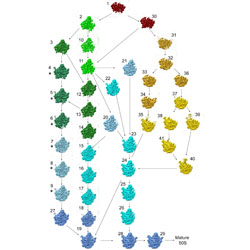
Quantitative mining of compositional heterogeneity in cryo-EM datasets of ribosome assembly intermediates
A workflow is presented that quantitatively mines cryo-EM data to isolate sets of structures with maximal diversity.
PubMed PMID: 34990602
PubMedCentral PMCID: PMC9891661
Structural basis of HIV inhibition by L-nucleosides: Opportunities for drug development and repurposing
The structure, function and future uses of L-nucleoside inhibitors is reviewed.
PubMed PMID: 35218925
PubMedCentral PMCID: PMC9232924 (available on 2023-07-01)
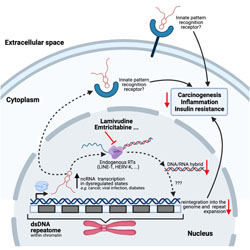
TRIM5α restriction of HIV-1-N74D viruses in lymphocytes is caused by a loss of cyclophilin A protection
A mutation in HIV-1 capsid blocks binding to cyclophilin A, leading to reduced infectivity of the virus.
PubMed PMID: 35215956
PubMedCentral PMCID: PMC8879423
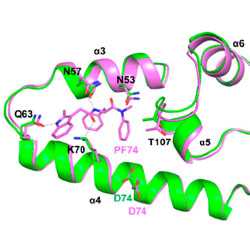
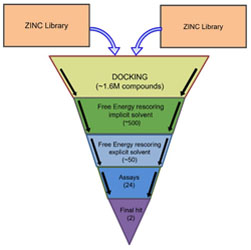
Structure-based virtual screening workflow to identify antivirals targeting HIV-1 capsid
A combination of computational docking and advanced free energy estimation is used to discover new HIV-1 inhibitors.
PubMed PMID: 35262811
PubMedCentral PMCID: PMC8904208
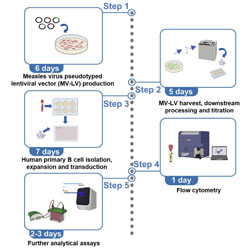
An optimized measles virus glycoprotein-pseudotyped lentiviral vector production system to promote efficient transduction of human primary B cells
A efficient protocol is presented to package complex transgene-containing lentiviral vectors.
PubMed PMID: 35284833
PubMedCentral PMCID: PMC8914380
Covariation of viral recombination with single nucleotide variants during virus evolution revealed by CoVaMa
Analysis of Flock House virus sequences reveals that recombination and point mutations of the genome are intimately linked.
PubMed PMID: 35018461
PubMedCentral PMCID: PMC9023271
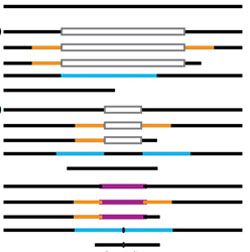
Prion-like low complexity regions enable avid virus-host interactions during HIV-1 infection
Several host cell proteins bind to HIV-1 capsids using unusual prion-like segments.
PubMed PMID: 36202818
PubMedCentral PMCID: PMC9537594
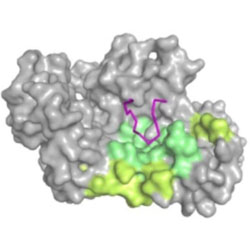
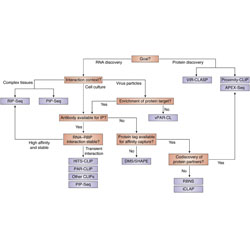
Next-generation sequencing: A new avenue to understand viral RNA-protein interactions
Recent advances in next-generation sequencing technologies have allowed the characterization of viral RNA-protein interactions.
PubMed PMID: 35413291
PubMedCentral PMCID: PMC8994257