B-HIVE Structure Highlight: Targeting 5′ leader RNA variations
Breaking down the 5’ leader
The HIV-1 5’ leader is a key repository of RNA-based instructions for viral life cycle processes. Because these processes include viral translation and packaging of genomes into new viral copies, it’s important to understand how mutations in this region affect viral function. Knowing what breaks the activity of the 5’ leader could lead to new therapeutic mechanisms.
At the Scripps Research Institute, the lab led by B-HIVE Collaborative Development Program grantee Megan Ken is exploring how mutations in the 5’ leader result in downstream effects of RNA function in cells. Based on previous work that correlated protein binding to a single RNA in the 5’ leader with activity of the RNA in that cell, Ken and her colleagues are developing high-throughput assays to explore other mutations consequential for viral function.
Expanding this effort is not without its challenges. The team hopes to evaluate tens of thousands of mutations throughout the 5’ leader, which involves many potential binding proteins. Current assays can evaluate a single mutant in a single cell at a time. Ken’s lab is developing a method to make stable cell lines, with thousands of transfected mutants, and measure viral cell activity all at once.
Interdisciplinary combinations
Suggestions, shared resources, and protocols from other B-HIVE members and Scripps Research Institute colleagues have been key to developing the stable cell lines. B-HIVE investigators Karin Musier-Forsyth and Wei-Shau Hu, in particular, have been instrumental in designing and pursuing these studies. Drawing on their experience, Ken says, has helped her lab create something new.
Her lab is set up to take full advantage of experimental and computational expertise on her research team. Computational researchers are set up just outside of the wet lab, in close proximity to experimental research. Both groups inform the work of the other: experimental work shapes computational simulations, which in turn shape the designs of new experiments.
How this lab is advancing HIV research
With enough high-quality assay data, Ken hopes to train machine learning models to understand the relationship between RNA activity and mutation sequence. Once trained, a model may be able to then predict downstream consequences of certain RNA mutations. A successful model can provide valuable insight about viral function and help evaluate new mutants observed by clinicians. An unsuccessful model can likewise yield valuable information, as researchers work to understand the basic biological mechanisms that the model failed to capture.
Ken also hopes that a high volume of data will identify RNA mutations that render the virus non-functional. An understanding of the structural factors behind that non-functionality may lead to development of small molecules that can reduce functionality in the wild-type virus. Pairing NMR structural data with computational and experimental screening hopefully will accelerate the drug discovery process.
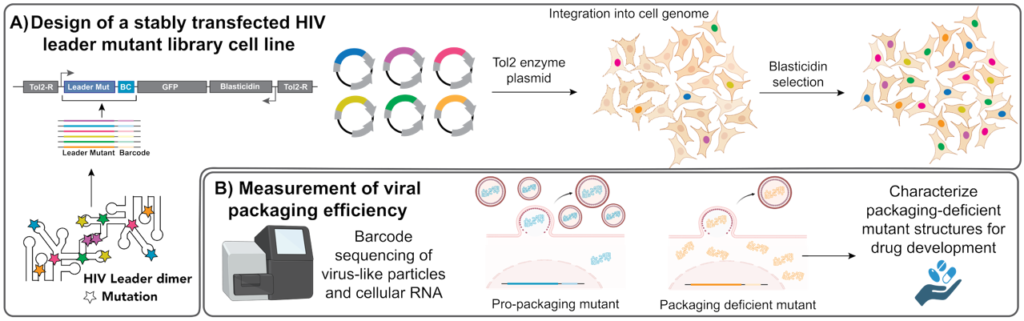
A) Design concept for our high-throughput viral packaging assay. Our mutational library contains 10,000 mutations of the HIV 5’ leader, predicted to change the structure of the leader in cells. We use the Tol2 system to stably transfect the entire library into cells, such that there is only 1 mutant integrated per cell.
B) We then express viral proteins to package these sequences, and use next generation sequencing of both the virus-like particles and the cellular RNA to determine how well each mutant packages RNA. Mutants that cause packaging defects will be further studied as possible targets for antiviral drug development.
Meet the Researcher
How did you get interested in science?
I was always a really curious kid. I was generally interested in science, enough to go to Space Camp as a teenager, but because I came from a background of folks who didn’t have the opportunity to get a lot of education, I didn’t understand when I was growing up that doing research was a possible career path.
What I did know was that I really loved my AP biology class. I had a great teacher, Dawn Norton, who really got me excited about biology. I thought that the only way to continue learning about biology was by going into medicine, so I went to Washington University in Saint Louis for college, intending to become an MD. Before I began, I got an email about applying to an HHMI-funded summer research experience, where you would come to campus the summer before freshman year and do a research internship. I was like, “Aah! that’s so cool. I’ve never heard of that. I don’t even know what that is!” To this day, I don’t understand how I was accepted, but by some miracle, I was. From the first day of that program, I was hooked on scientific research and continued throughout my undergraduate career.
A huge part of what made my undergraduate research experience so impactful was that I had an amazing research advisor, Audrey Odom John. When I began in the lab, I still didn’t understand how research worked, or how a person can create new knowledge. Up until this point, everything I learned was very prescriptive. “This is how this works, this is how that works. Memorize it.” And she very quickly flipped that script for me, instead focusing on questions – how to define them and then design experiments that aim to not only answer them, but lead to more questions. I loved every second of it and by the time I graduated she convinced me to do an MD-PhD instead of focusing solely on clinical training. I followed in her footsteps and did my MD/PhD at Duke University, and am very grateful for all of the excellent mentors I had there, both in my scientific and medical training. But by the end of my time in the program, I realized that practicing clinical medicine didn’t really make sense for me anymore. My heart was in basic research and I had already learned all the clinical knowledge needed to realize my scientific goals. So that’s why I decided to start my lab instead of going to residency.
Tell us about the lab where you did this work.
I’m in an interesting program called the Scripps Fellows. It’s a program that promotes early independence by allowing students who are right out of their PhD, or within one year of doing their postdoc, to start setting up their labs without doing a traditional postdoc research experience. Scripps has many effective supports in place to help you be successful. The idea is to provide you with enough funds to purchase reagents and hire scientists to be in your lab and then pair you with a senior faculty sponsor who can provide access to lab space, equipment, and, of course, excellent mentorship.
That person for me is Jamie Williamson, who’s also in B-HIVE, which was my entree into this group of people. He and his lab members have been very gracious; my lab is currently contained literally within his lab. We’ve formed this symbiotic relationship that allows us to work alongside one another in physical space, but also has facilitated really exciting research collaborations. I have recently been promoted to assistant professor, which will come with my own space, but we look forward to continuing close collaboration with Jamie regardless.
Research is ultimately about people, and I feel extremely fortunate that folks in my lab are so awesome. I have three graduate students through the Scripps graduate program, the first of whom agreed to rotate before I even moved to San Diego and joined in 2024. She’s the scientist driving this high-throughput viral packaging project. I have two postdocs who are doing purely computational work on developing highly accurate structural ensembles of RNA and docking RNA with small molecules for drug targeting. That’s in collaboration with Stefano Forli, who’s another B-HIVE member at Scripps. And then we also have several technicians who’ve been moving a lot of collaborations forward and generating important preliminary data. Everyone is really kind, curious, and engaged in each other’s work, so it’s a really fun group to solve problems with.
What were the biggest challenges with this study?
The biggest challenge with the study is really the challenge of starting a new lab. We are largely doing experiments that I’ve done before, but setting them up from scratch is quite different from learning something in an already established lab. I came from a large lab that had all of these things working, and the challenge is coming to a new place and make them work again, and then also set up new experiments that I’ve never done before. For example, I had never done a viral packaging experiment myself before I came here. Communicating and collaborating with experts, like B-HIVE investigators, has been instrumental in our success so far.
Because we’re so interdisciplinary, there are a lot of things we need to get started at once. We need to be booting up computational programs and software pipelines. We need to get several types of assays working both in vitro and in cells. We need to get NMR experiments running on the new Scripps spectrometers. There are a lot of things that have to be built separately and then come together farther down the line. Practically speaking, that’s the biggest challenge.
From a scientific perspective, the biggest challenge with this high-throughput method is being able to find a way where you can test many mutants at once, but by minimally disrupting the native context of these viruses. You have to be thinking about the data you will get at the end and how you will interpret it when you design it from the beginning. When you get the results, you will always ask yourself: Is this based on something real or is the nature of doing the experiment in high-throughput fundamentally altering the results? We have done our best in the design phase to control for certain aspects of our interventions, but of course we are going to have to do a lot of downstream validation of the results to make sure that what we’re seeing is real.
What are you working on now?
A lot of the work up to this point has been establishing the high-throughput assays and building tools to design the mutant libraries that will give us insights into viral function. We are now executing those experiments and hope to share our results very soon. Another portion of my B-HIVE proposal is to look at the co-transcriptional folding pathway of the HIV leader RNA and understand not only the energetics of that landscape but identify potential intermediates that would make good drug target candidates. We have implemented a novel method developed by external collaborators that should help us understand how the kinetics of co-transcriptional folding allows HIV to have a diversity of structure-function relationships within the same sequence. Ultimately, we hope to apply virtual screening techniques being optimized by our computational scientists, in collaboration with the Forli group, to target conformations identified by our high-throughput viral packaging assays and co-transcriptional folding assays.

